Metagenomics workflow for hybrid assembly, differential coverage binning, metatranscriptomics and pathway analysis (MUFFIN) | PLOS Computational Biology

Ten Years of Maintaining and Expanding a Microbial Genome and Metagenome Analysis System: Trends in Microbiology
![PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/88cd5fdbfb2dba16a6b23987d30ad9437ff0c805/9-Table3-1.png)
PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar

Genome-Resolved Metagenomics Is Essential for Unlocking the Microbial Black Box of the Soil: Trends in Microbiology

HiFi metagenomic sequencing enables assembly of accurate and complete genomes from human gut microbiota | Nature Communications

MetaRef comprises automated processing of available microbial genomic... | Download Scientific Diagram
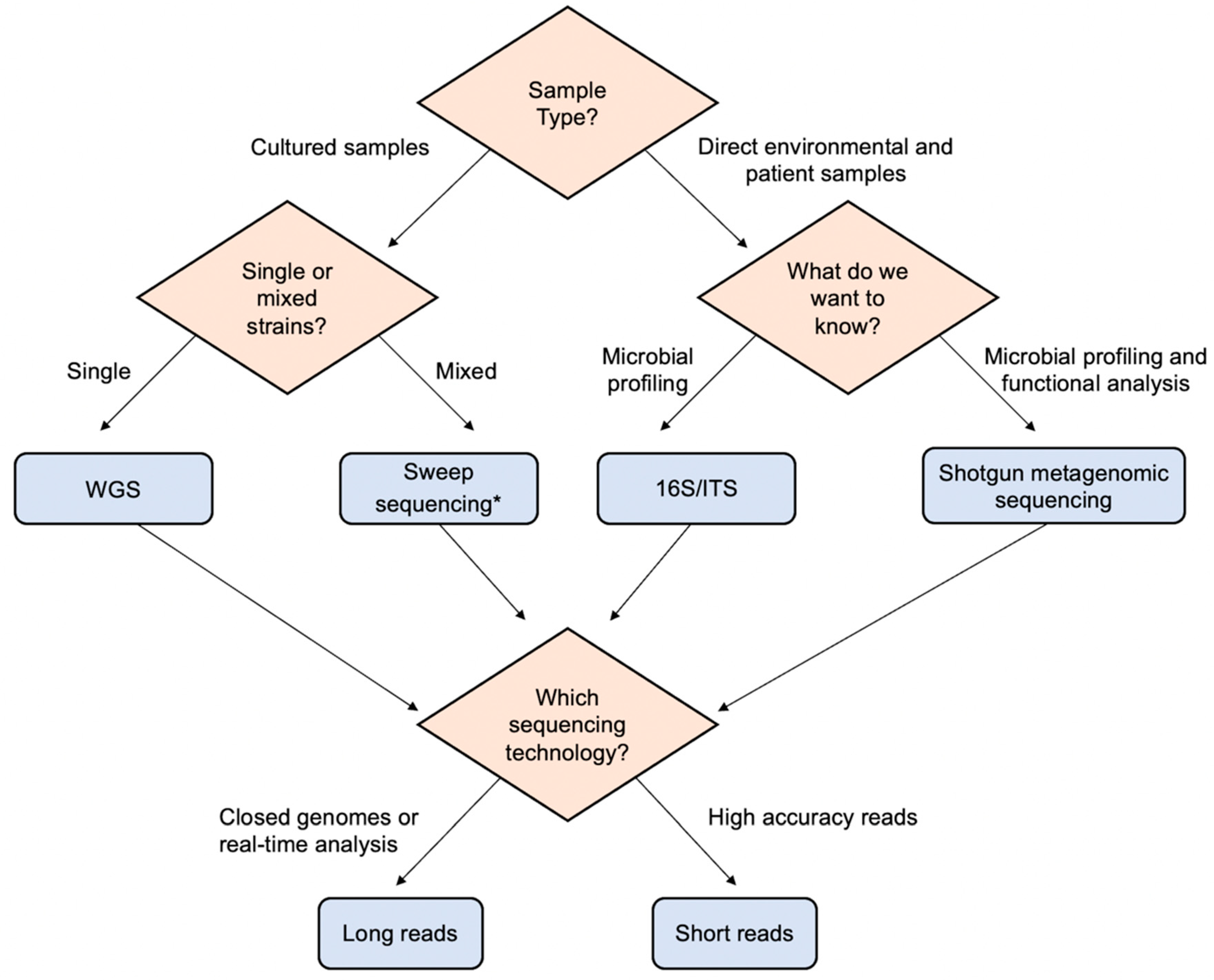
IJMS | Free Full-Text | Combination of Whole Genome Sequencing and Metagenomics for Microbiological Diagnostics
![PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/88cd5fdbfb2dba16a6b23987d30ad9437ff0c805/14-Figure3-1.png)
PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar

Assembly of 913 microbial genomes from metagenomic sequencing of the cow rumen | Nature Communications

Recovering metagenome-assembled genomes from shotgun metagenomic sequencing data: Methods, applications, challenges, and opportunities - ScienceDirect
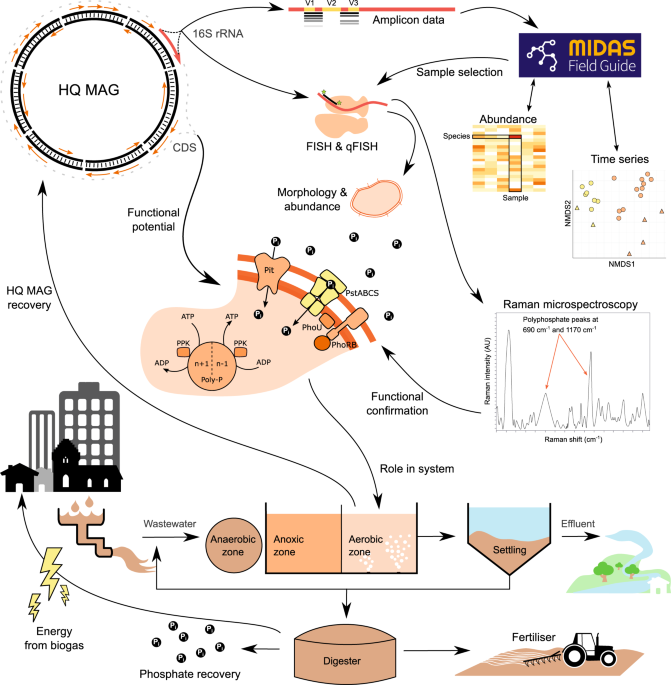
Connecting structure to function with the recovery of over 1000 high-quality metagenome-assembled genomes from activated sludge using long-read sequencing | Nature Communications
![PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/88cd5fdbfb2dba16a6b23987d30ad9437ff0c805/9-Table2-1.png)
PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar
![PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/88cd5fdbfb2dba16a6b23987d30ad9437ff0c805/13-Figure2-1.png)
PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar
![PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/88cd5fdbfb2dba16a6b23987d30ad9437ff0c805/12-Figure1-1.png)
PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar
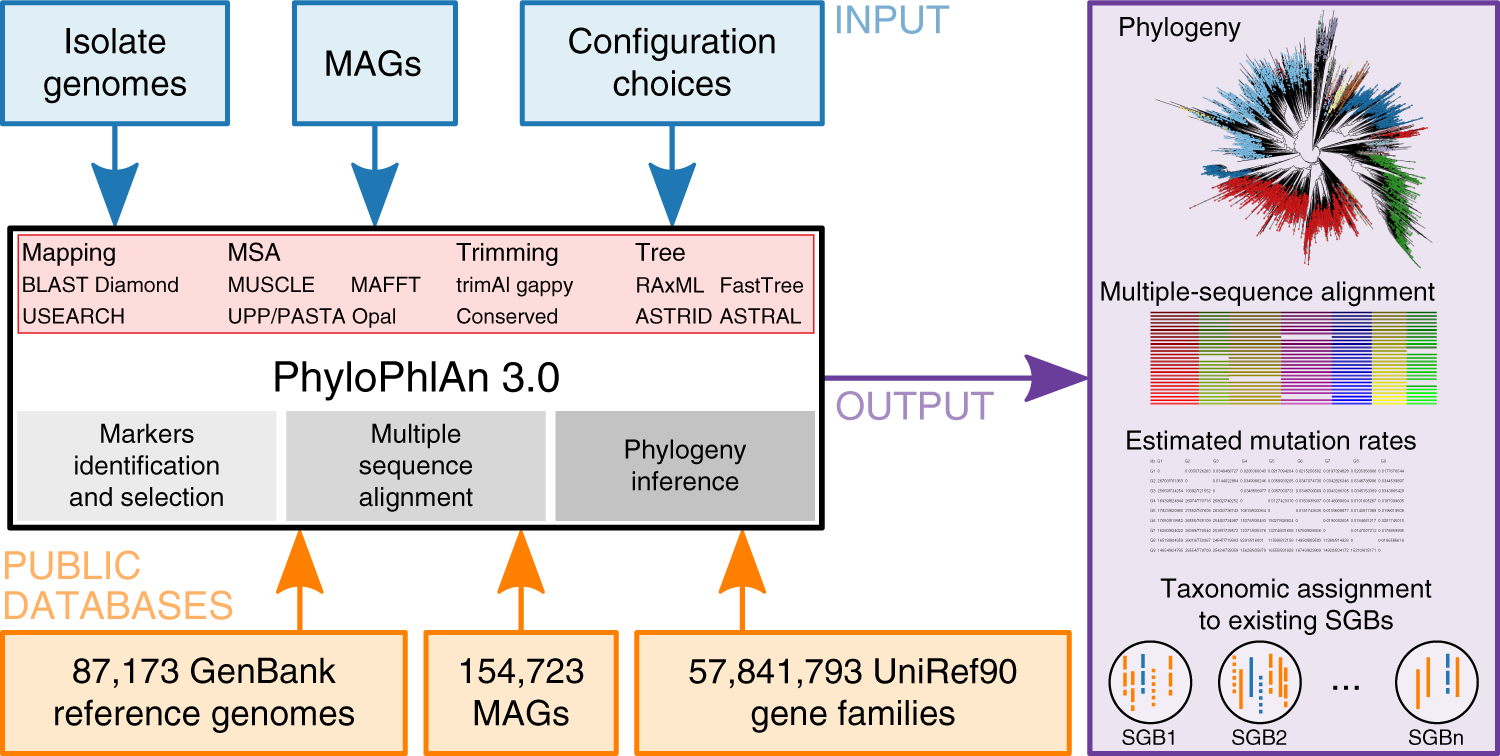
Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0 | Nature Communications
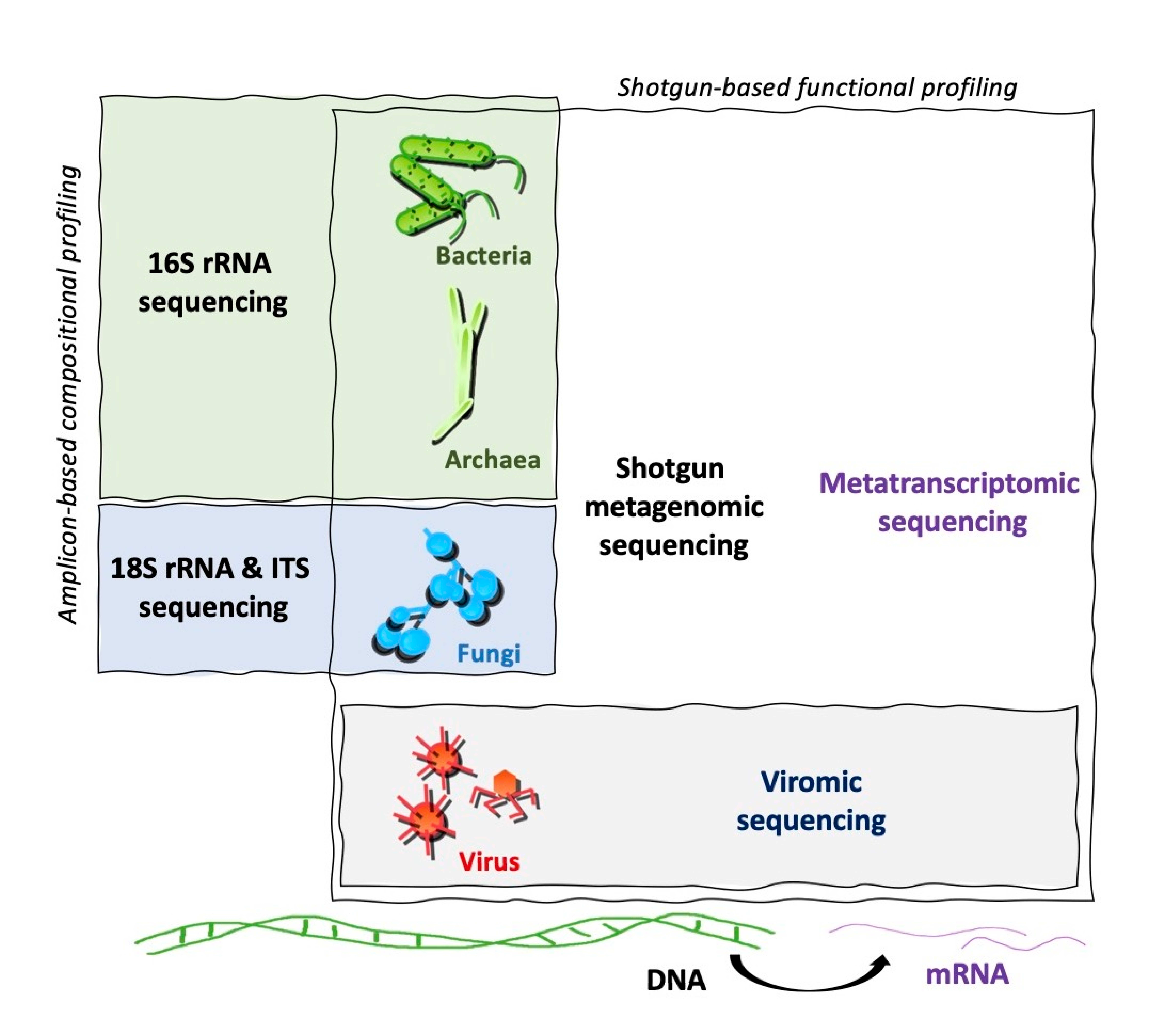
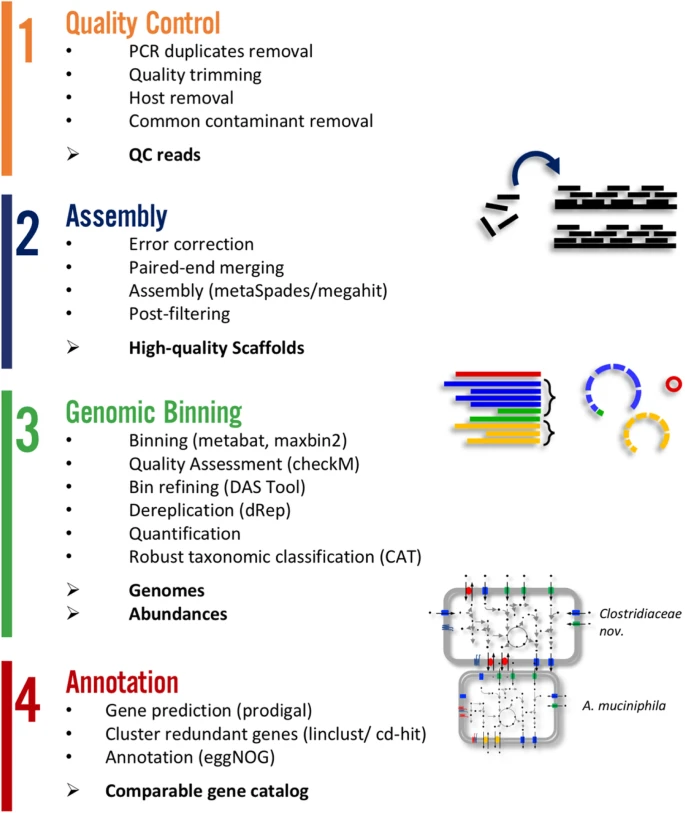
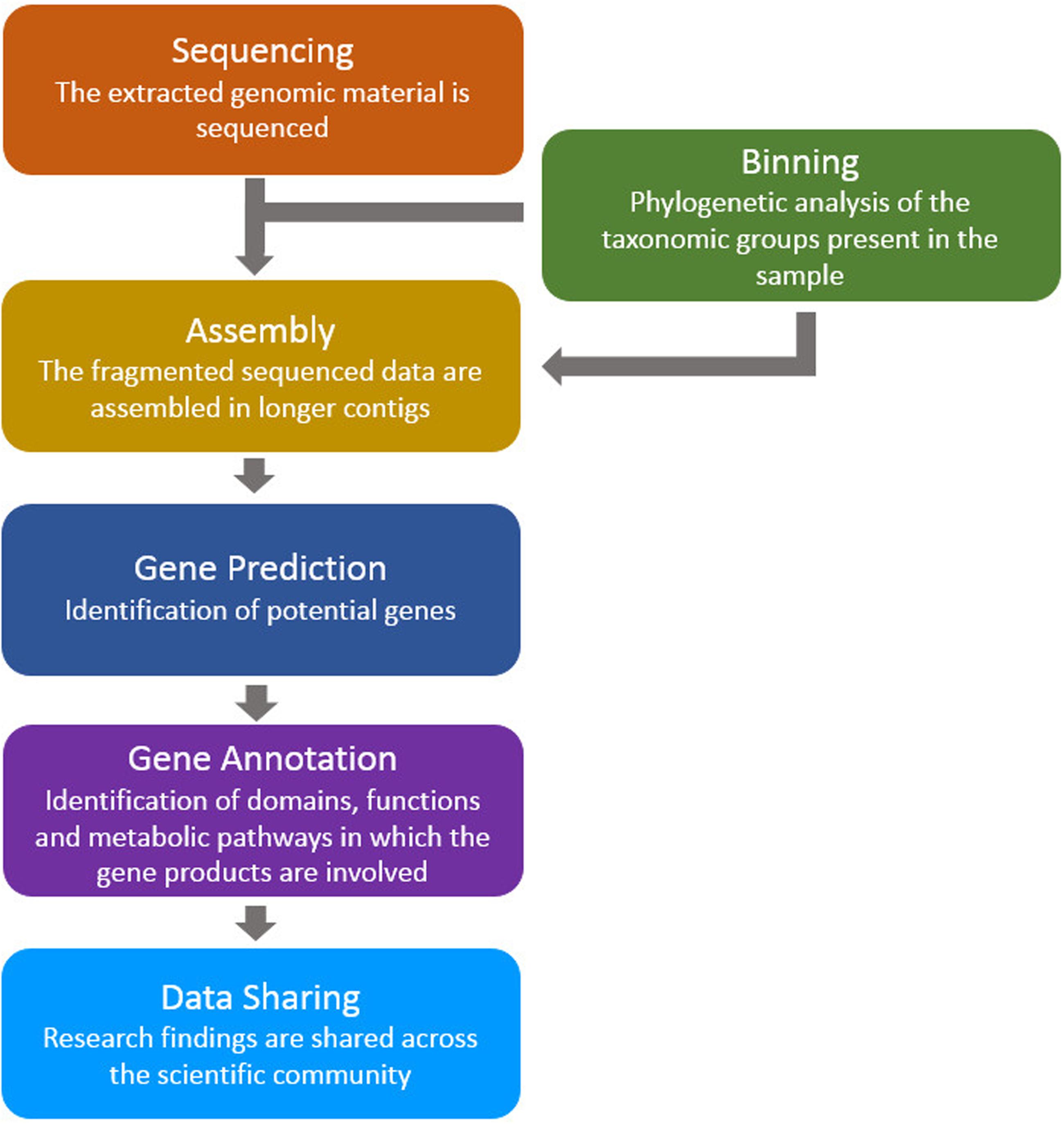
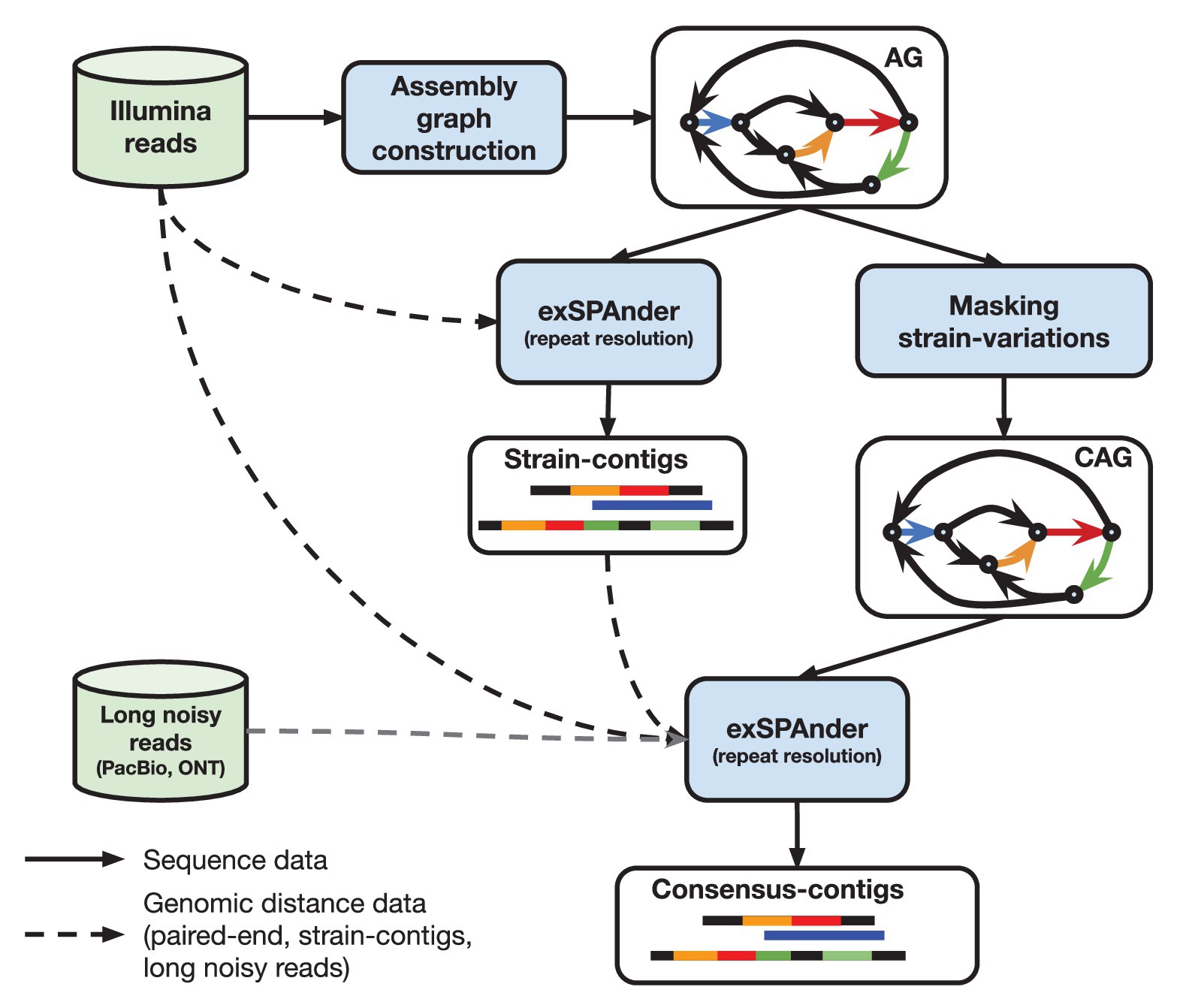
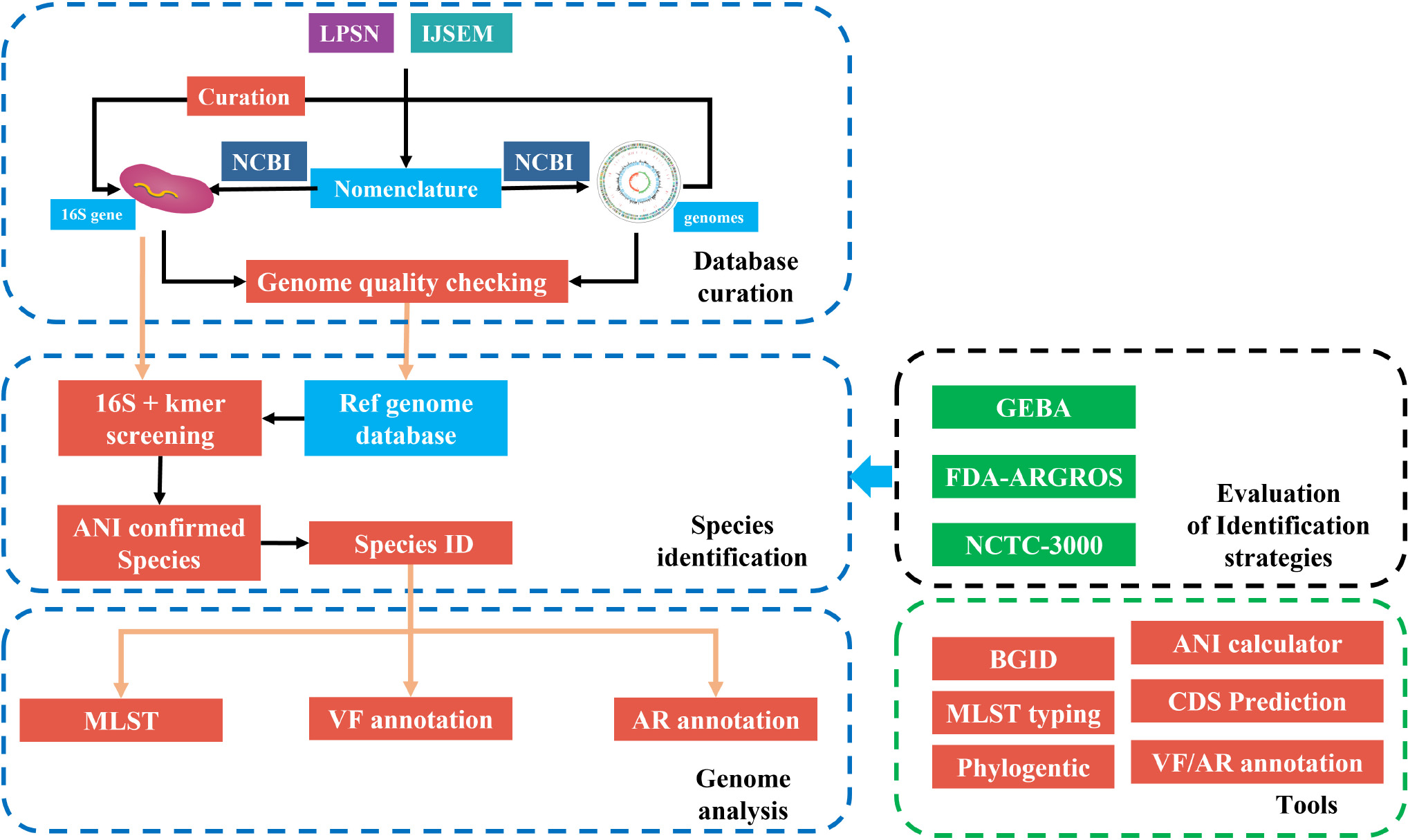

![DATMA: Distributed AuTomatic Metagenomic Assembly and annotation framework [PeerJ] DATMA: Distributed AuTomatic Metagenomic Assembly and annotation framework [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/9762/1/fig-1-full.png)
